An online repository of AmpliconArchitect outputs. Explore and download focal amplification predictions and annotations.
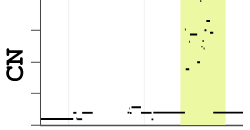
Copy-number data exploration
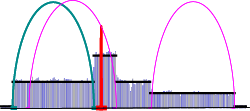
AmpliconArchitect output files

Focal amplification classifications
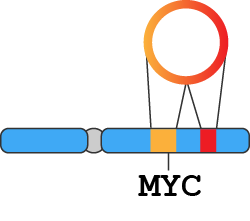
Amplicon coordinates and gene contents
| Name | Description | Last Updated | Sample count | |
|---|---|---|---|---|
| Featured | Hartwig Medical Foundation - HMF | Collection of metastatic tumor samples from HMF. Included in Kim et al., 2024. Reclassified with AC… | 2026-04-15T22:12:28.064719 | 4,170 |
| Featured | PCAWG | Samples processed by Verhaak and Kim labs, filtered for coverage, duplicates and other criteria as … | 2026-04-15T22:03:57.544610 | 2,095 |
| Featured | CUGA | Chinese Urothelial Carcinoma Genome Atlas (CUGA) samples processed by DREAM lab (Lv, Zeng, and Li e… | 2025-03-11T01:56:16.364926 | 1,411 |
| Featured | CCLE | AA results from CCLE with metadata, updated for AC version 1.4.2 | 2026-04-04T21:30:05.910142 | 329 |
| Featured | Pediatric Pancancer | Tumor samples from St. Jude and PBTA | 2026-03-26T20:55:46.969563 | 3,796 |
| Public | GBM39 | GBM39 (FF-4, naive) from Turner 2017 | 2026-04-16T20:54:49.333399 | 1 |
| Public | COLO320 DM & HSR hg38 | GRCh38-based results for COLO320DM and COLO320HSR using sequencing data published in Wu et al., Nat… | 2024-02-20T22:16:22.120064 | 2 |
| Public | Medulloblastoma GRCh37 subset | Medulloblastomas from ICGC published in Chapman et al. 2023 | 2023-11-01T22:34:24.516799 | 275 |
| Public | 20251017_testing_SRR28999682_import_form_GenePattern | SRR28999682_20251017 | 2025-10-21T04:35:33.413824 | 1 |
| Public | TCGA unfiltered | TCGA samples processed by Verhaak and Kim labs, NOT filtered for coverage, duplicates and other cri… | 2026-04-13T19:18:26.421741 | 2,471 |
| Public | Nakagawa human-viral HPV analysis | Oropharyngeal cancer cell lines, including some HPV-negative cell lines. | 2024-11-20T01:04:13.378203 | 16 |
| Public | TCGA | Overlapping samples with PCAWG assigned to PCAWG project. Samples processed by Verhaak and Kim labs… | 2026-04-15T22:05:39.542427 | 101 |
| Public | PCAWG unfiltered | PCAWG samples processed by Verhaak and Kim labs, NOT filtered for coverage, duplicates and other cr… | 2026-04-13T23:53:36.442554 | 2,908 |
| Public | TR14 | TR14 cell line from Hung et al., 2021, re-analyzed with hg38 (thanks to Kaiyuan Zhu) | 2025-02-24T00:52:31.986595 | 1 |
| Public | Pradella et al. engineered murine Myc ecDNA | Induced Myc ecDNA and control samples with timecourse sampling from Ventura lab | 2024-03-27T19:08:31.477663 | 12 |
| Public | Medulloblastoma hg38 subset | Medulloblatomas from CBTN, St. Jude Cloud, and Rady Children's hospital, published in Chapman et al… | 2023-11-01T22:34:44.496681 | 208 |
| Public | COLO320DM | Low coverage hg19 COLO320DM used in Hung et al., Nature 2021 (ecDNA hubs) | 2023-05-08T17:43:50.493387 | 1 |
| Public | ec3D | AA outputs for cell lines analyzed in ec3D manuscript (Chowdhury, Zhu, Li, et al.) Interactive 3D v… | 2025-06-05T19:25:47.131745 | 7 |
| Public | Pradella et al. engineered murine Mdm2 ecDNA | Mdm2 mouse tumors. These were obtained by injecting subcutaneously Ras-infected ecMdm2 mouse fibrob… | 2024-03-27T19:09:40.130576 | 5 |
| Public | 20250728-test | 20250728-test | 2025-07-29T07:08:52.532202 | 8 |
| Public | HCC827 | Sample Preparation and Sequencing Protocol For the HCC827_ER (elotinib-resistant) and HCC827_DR (d… | 2024-11-20T20:03:14.874416 | 4 |
| Public | Prostate cancer cell and colorectal cancer cell line | SRR28999683 SRR28999682 SRR8236746 SRR9157088 (fastq input) | 2025-07-30T02:06:57.684511 | 4 |
| Public | ecDNA replication - Tang, Weiser, Wang, et al. 2024 | AA outputs for samples used in analysis of ecDNA replication differences between DM and HSR forms. | 2024-11-11T18:59:13.611683 | 6 |
| Public | 20250716_PC3_DM/HSR_COLO320DM/HSR | 20250716_PC3_DM/HSR_COLO320DM/HSR | 2025-07-29T07:02:53.528547 | 4 |
| Public | GLASS | GLASS (Glioma Longitudinal AnalySiS) samples processed by Verhaak and Kim labs, filtered for covera… | 2026-04-14T20:13:32.854434 | 140 |
| Public | Turner 2017 cell lines | Re-analyzed for hg38. Project contains some replicate cell lines. Check metadata for more info. | 2026-04-02T18:54:50.260374 | 162 |
News & Updates
- Leveraging large-scale genomic datasets to track extrachromosomal DNA across cancer stages
- Study of ecDNA in medulloblastoma reveals intratumor heterogeneity of ecDNA
- Study of ecDNA in Barrett's esophagus patients shows ecDNA arises early in cancer development
- Team eDyNAmiC receives $25 million grant to study the role of ecDNA in cancer
Luebeck J, Huang E, et al. "AmpliconSuite: an end-to-end workflow for analyzing focal amplifications in cancer genomes." bioRxiv. 2024.
To cite results obtained from the site, please refer to the uploader's associated publication where possible.
To cite results obtained from the site, please refer to the uploader's associated publication where possible.
What's publicly available?
| Projects | 25 |
| Samples | 18,137 |
Focal amplification counts
| ecDNA | 5,375 |
| BFB | 2,272 |
| Other fsCNA | 15,905 |