Genomic patterns of hybridisation and its role in ecological adaptation
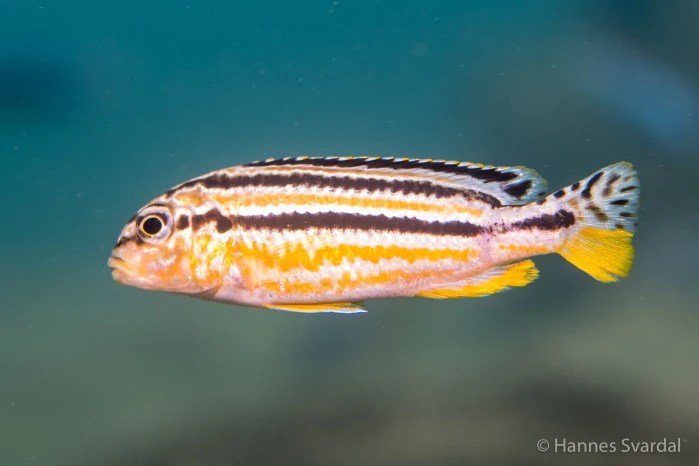


Recent genome studies suggest that hybridisation and genetic exchange among closely related species is more common than previously thought. Consequently, a central question in the study of biodiversity is the effect of genetic exchange on the formation and maintenance of species diversity. With more than 800 closely related species, the Lake Malawi cichlid fish adaptive radiation provides an intriguing model to study the frequency and evolutionary role of interspecific genetic exchange. Malinsky et al. (2018) found strong evidence for extensive gene flow early on in the Malawi radiation and made some links to adaptation. However, due to limitations in sampling and statistical inference methods, we are still lacking a comprehensive picture of the abundance and evolutionary role of genetic exchange in Malawi cichlids and in most other organisms. In this project I will establish a genomic framework for the joint inference of species relationships and genetic exchange, applying it to a unique dataset of more than 2000 genomes from 276 Malawi cichlid species to gain unprecedented insight into the abundance of genetic exchange between different populations, species and genera. Furthermore, I will refine a statistical method I developed during my Master’s to test whether selection has acted on exchanged genetic material. In summary, this project will yield widely applicable genomic tools and insight into the role of gene flow in one of the most intriguing vertebrate radiations.
Team: Sophie Gresham, Hannes Svardal
Collaborators: Richard Durbin (University of Cambridge), George Turner (Bangor University), Milan Malinsky (University of Bern)
Funding: FWO PhD fellowship
The genomic basis of variation in body armour in a girdled lizard



Osteoderms are bony elements that are expressed in the skin of a few disparate groups of tetrapods (i.e. in crocodiles, turtles, armadillos, and some lizard and frog species) – but not in other taxa. In humans, osteoderms are frequent complications of injury and in a few rare inherited disorders. Osteoderms spark interest because they are ecologically relevant (they are likely to function in body protection, thermoregulation and water budget maintenance, in mineral storage) but at the same time exhibit an unusually binary distribution (i.e., they are expressed completely, or not at all). The latter element facilitates research into the genomic substrate of the trait. One species of cordylid lizard, Hemicordylus capensis, uniquely displays intraspecific variation in osteoderms: the trait has evolved repeatedly and therefore is present in some populations, but not in others. The species thus offers exceptional opportunities for learning how, why and when this remarkable trait evolves. With this project, we aim to resolve those issues through a thoroughly integrated approach combining state-of-the-art genomic, functional morphological and ecological techniques. We will also explore if we can extrapolate the findings on this study system to other taxa that (occasionally) express osteoderms, including humans. The project will allow a rare complete view of the evolution of an ecologically relevant phenotypic characteristic with a remarkably discontinuous variation and an unusually disparate taxonomic distribution.
Team: Genevieve Diedericks, Henrique Leitão, Erik Matthysen, Kris Laukens, Hannes Svardal, Raoul Van Damme
Collaborators: Simon Baeckens, Chris Broeckhoven, Cang Hui (University of Stellenbosch), Anton du Plessis (University of Stellenbosch)
Funding: BOF-GOA, FWO PhD fellowship
The molecular and phenotypic basis of adaptation to heavy fishing
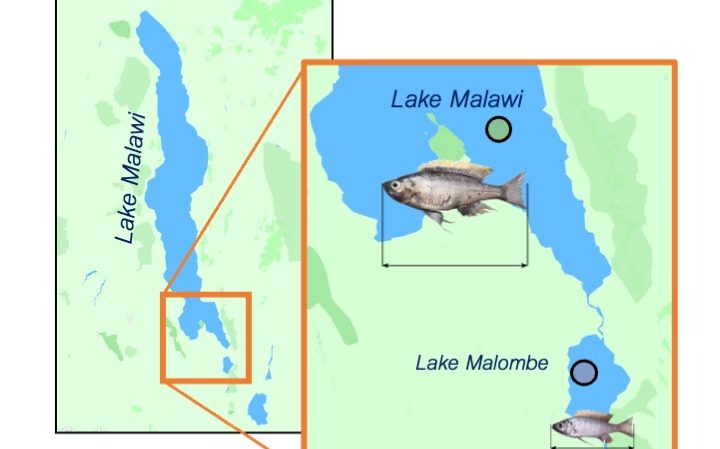

We currently lack a detailed understanding of how organisms rapidly adapt to environmental changes, which is key in evaluating and predicting human impact on nature, uncovering the genetic basis of adaptive traits, and gaining insight into fundamental evolutionary processes. Evolutionary response to a direct form of human impact, fishing,has often been discussed but evidence from natural systems is scarce. To address this, I will investigate genetic and phenotypic changes in Malawi cichlid fish following ~40 years of intense fishing. In particular, I will address life-history trait changes. Genome sequencing of museum specimens collected before and during fishing will give unprecedented insight into genes under selection. Extensive genomic resources available for Lake Malawi cichlids will allow me to investigate the evolutionary history of genes used in recent adaptation. I will leverage the ease of breeding cichlids in the lab to experimentally quantify genetic and environmental differences in traits implicated in fisheries-induced evolution. Furthermore, I will use state of the art (ancient) DNA and RNA sequencing technologies and bioinformatic methods to identify the genomic signature & molecular pathways involved in rapid life history adaptation. The combination of genome sequencing and controlled breeding experiments will greatly advance our understanding of how genomes can rapidly adapt to fishing and the link between selective pressures, phenotypes and genotypes.
Team: Alex Hooft, Wilson Sawasawa, Gudrun de Boeck, Hannes Svardal
Collaborators: Ludovic Orlando (CNRS)
Funding: BOF, FWO PhD fellowship
The role of chromosomal inversions in the rapid evolution of biodiversity

Understanding how biodiversity evolves is a defining question in evolutionary biology and instrumental in conservation and mitigation in the face of global change. Recent work, including ours, suggests that new combinations of old genetic variants are a driving force in rapid adaptive diversification. There is a large body of theory predicting that inversions – chromosomal rearrangements in which a segment of a chromosome is reversed end to end – play a decisive role in this process, for example, by linking together adapted alleles. There is strong evidence for the role of inversions in adaptation of natural species, but we are lacking a quantitative assessment of the role of inversions in diversification of a large group of related species. We will fill this gap by characterising the occurrence, evolution, and role of inversions across the fishes of the extraordinarily diverse Lake Malawi cichlid adaptive radiation. For this, we will combine innovative molecular and computational approaches with a unique set of hybrid crosses between species up to two million years divergent.
Team: Jaysmita Saha, Hannes Svardal
Collaborators: Luka Moritz Blumer (University of Cambridge), Richard Durbin (University of Cambridge)
Funding: BOF
Speciation in pebble beaches – Exploring an interstitial fish radiation
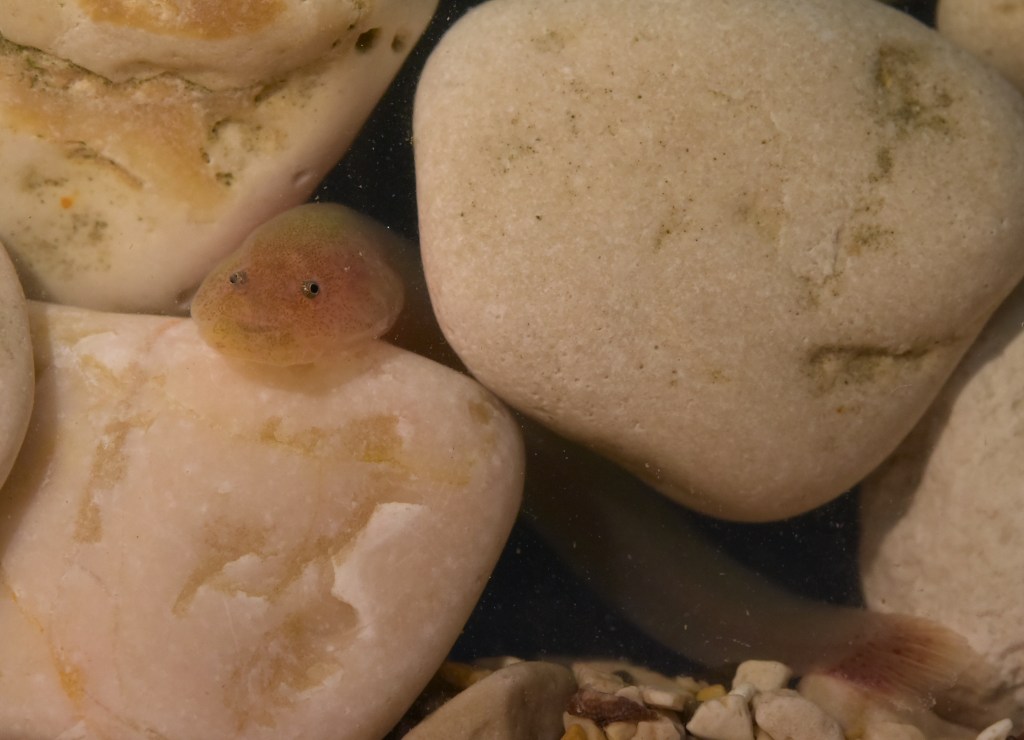
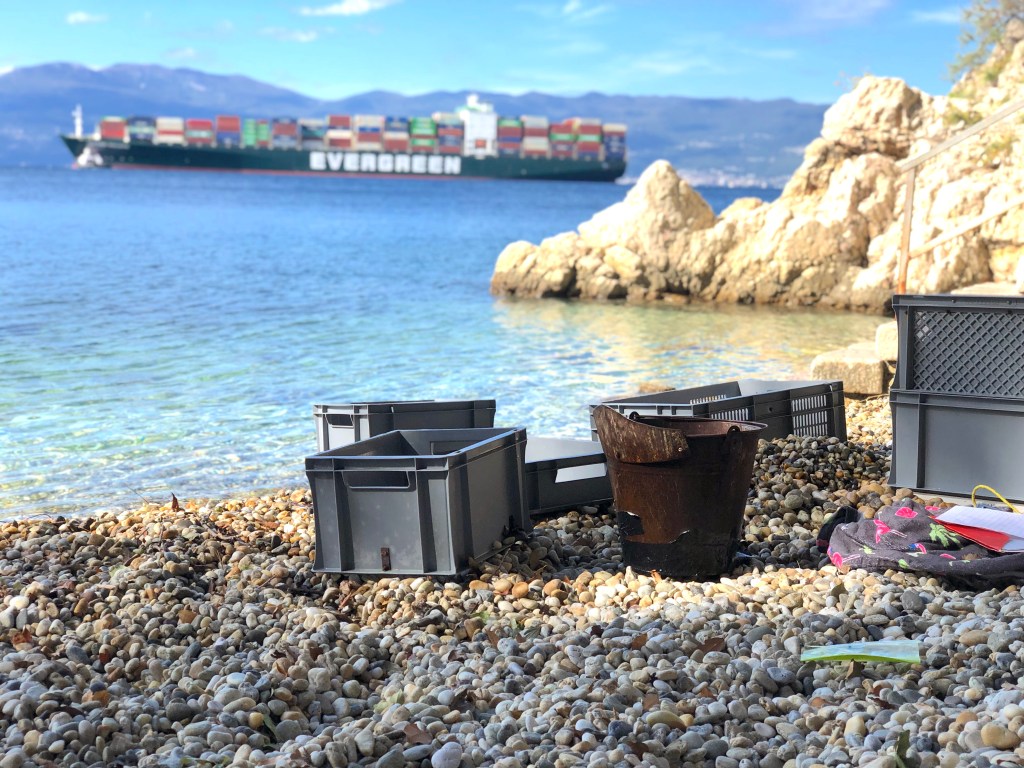
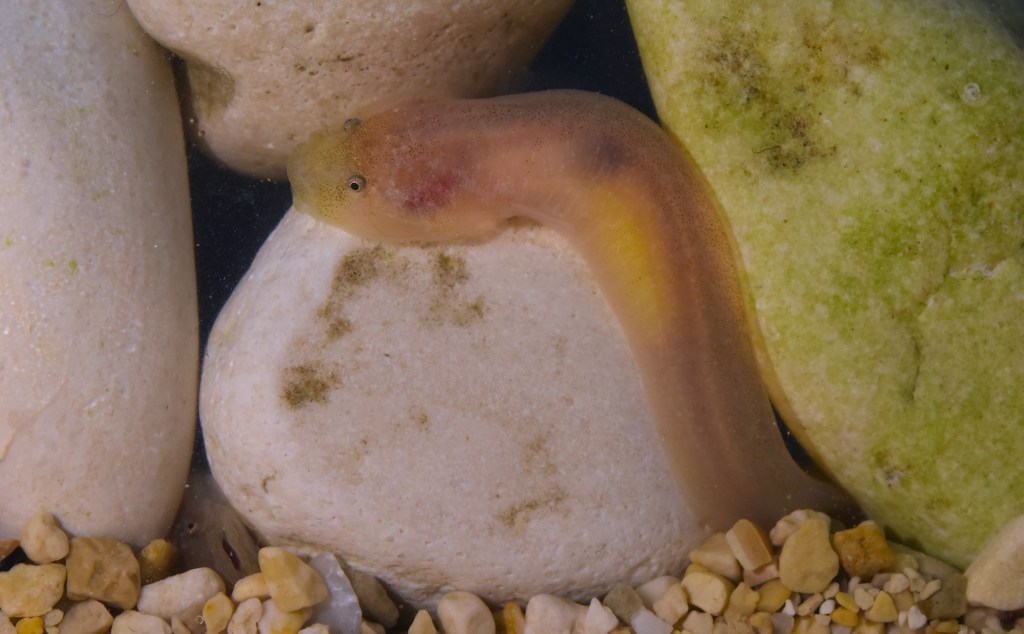



Mostly invisible to the human eye, miniaturised and characterized by a hidden lifestyle, “cryptobenthic” fishes are among the scientifically least studied groups of vertebrates on this planet. Denoted as the “hidden half”, cryptobenthic fish diversity is thought to be considerably underestimated, even in generally well-studied biomes like the Mediterranean Sea. This is especially true for the monotypic clingfish genus Gouania, a Mediterranean endemic, inhabiting the interstitial of intertidal gravel beaches. Indeed, only a few species of vertebrates worldwide cope with the life-hostile conditions prevailing in this particular environment. Preliminary investigations showed that the genus harbours at least five (four more than originally thought) fairly distinct species that have been diversifying for millions of years. Additionally, Gouania comes in two morphotypes, “slender” and “stout”, that convergently evolved in the eastern Mediterranean and the Adriatic Sea. This independent evolution of morphology likely mirrors the adaptation to certain microhabitats which was shaped by different selective pressures acting in these environments. Using whole genomic data, but also classical approaches, such as taxonomy, I aim to illuminate this enigmatic system from different methodological angles in the course of my ÖAW DOC fellowship project.
Team: Maximilian Wagner, Stephan Koblmüller (Univ. Graz), Hannes Svardal
Collaborators: Marcelo Kovacic (NhM Rijeka), Stamatis Zogaris (HCMR, Anavyssos), Richard Durbin (Univ., Cambridge), Kevin Conway (Texas A&M Univ.), Leon Hilgers (Museum Alexander König, Bonn), Gudrun de Boek (Univ. Antwerp)
Funding: ÖAD (Austrian Academy of Science) – DOC-Fellowship
Small-scale agroecological farming as a promoter of genetic diversity in insect pollinators and pests: Integrated ‘omic approaches for an African test case
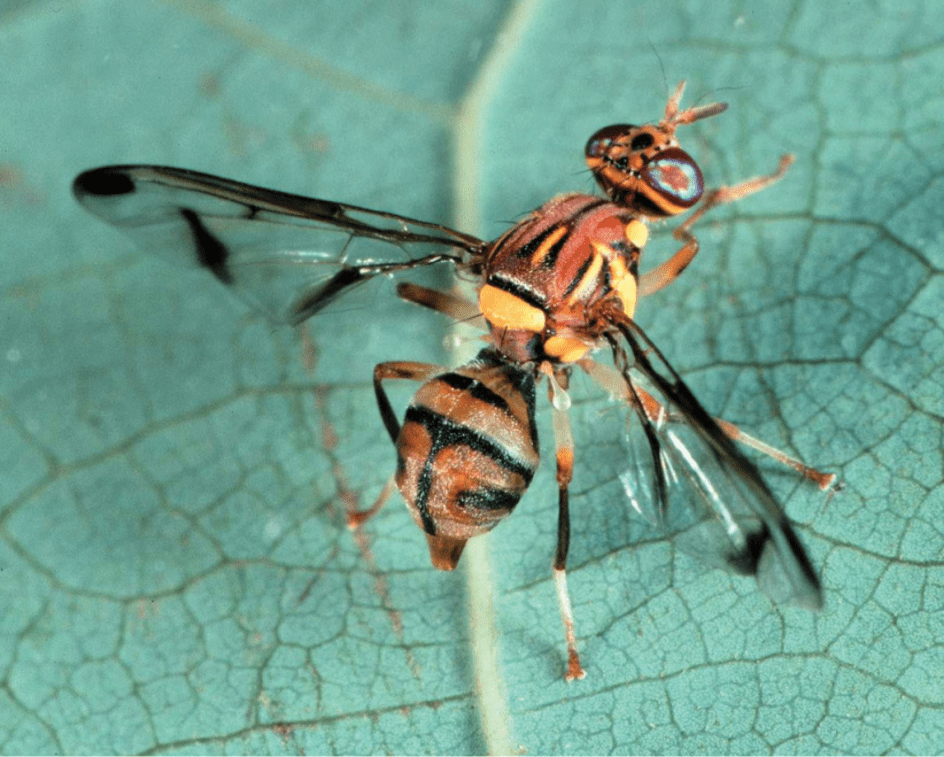



With the global decrease in pollinators and the increased awareness around the damages of pesticides (both to the environment as human health), there is an increased interest in finding better alternatives to current pesticide-intensive agricultural practices. In this project, we will focus on agroecological practices in small-scale farms in Morogoro, Tanzania, in collaboration with the Sokoine University of Agriculture. The overall BRAIN project will investigate the impact of agroecological farming on the yield, the economic feasibility of the practices, and the impact it has on the diversity and adaptive/plastic responses of both pollinators and pest species. We focus on two major functional groups of insects, namely hoverfly pollinators (Diptera, Syrphidae) and fruit fly pests (Diptera, Tephritidae) as targets and proxies for the evaluation of pollinator and pest biodiversity in contrasting agroecosystems. Different integrated omics approaches (metabarcoding, whole-genome sequencing, transcriptomics) will be explored to provide experimental support to the general paradigm of agroecology as a promoter of insect (genetic) diversity and relevant information on adaptive and plastic responses to the agroecological environment.
Team: Nele Mullens, Hannes Svardal
Collaborators: Massimiliano Virgilio (Royal Museum for Central Africa), Marc De Meyer (RMCA), Kurt Jordaens (RMCA), Nicolas Vereecken (Université Libre de Bruxelles), also in collaboration with the Royal Belgian Institute for Natural Sciences and the Sokoine University of Agriculture
Funding: BRAIN-BE 2.0
The use of alternative genomic markers to reconstruct the complex evolutionary history of neotropical small felids
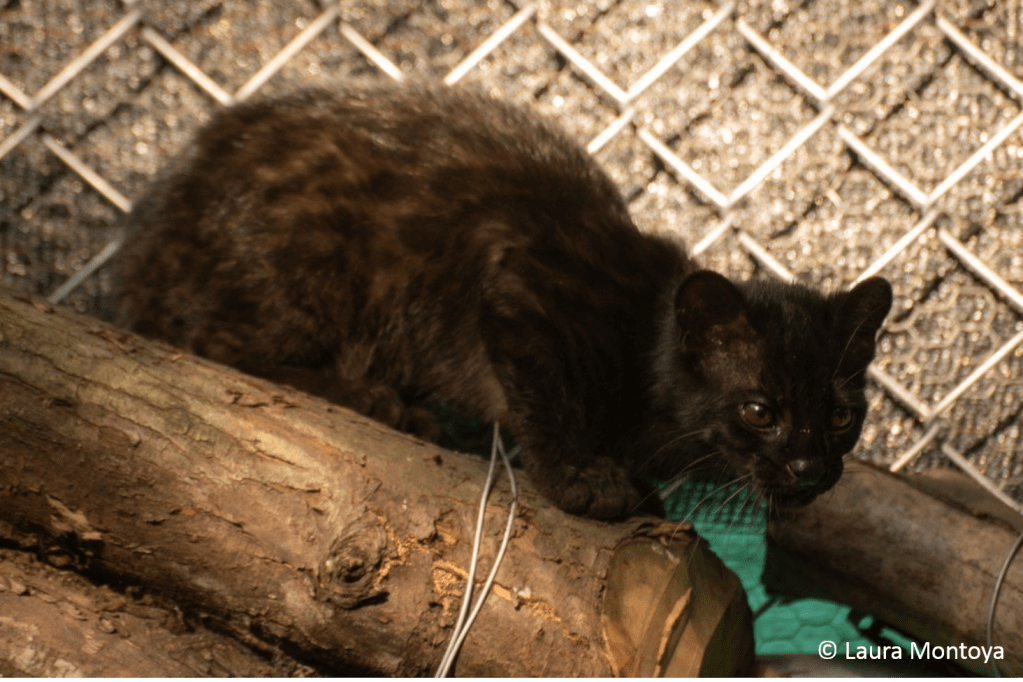
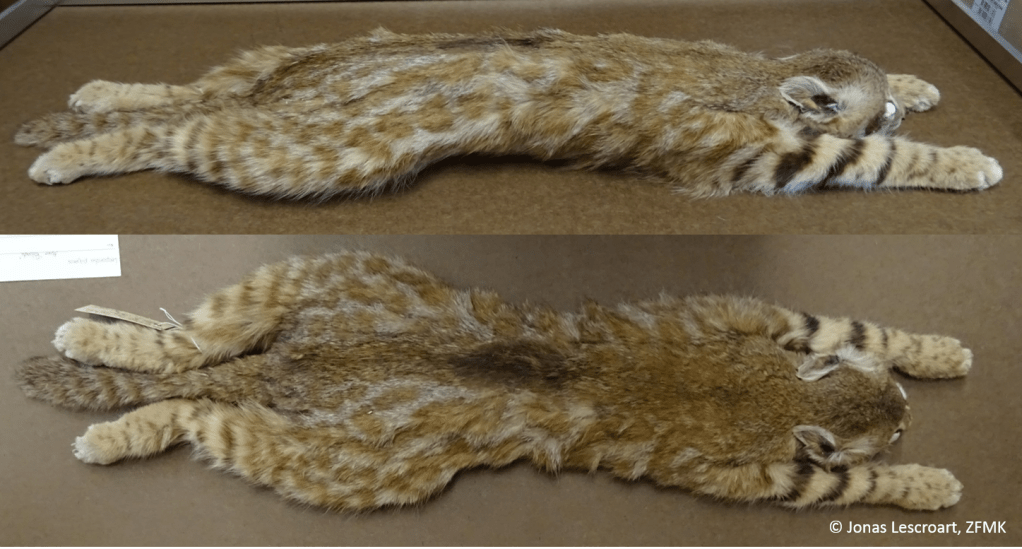
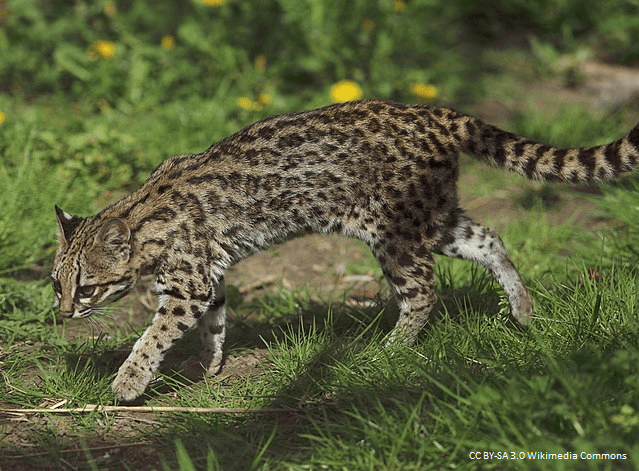



Tracing the evolutionary origin of species can be a challenging task, especially when these species cross-breed and thereby cloud the genetic record of their evolution. Fast-developing techniques in acquiring genetic information have improved to the point where complete genomes can be routinely sequenced. One of the upcoming questions is how to make optimal use of this massive amount of data. This research project combines the above challenges, by using complete genomes of a group of cross-breeding cat species from Latin America. The objective is to optimize the use of vast genomic data to elucidate evolutionary history in a complex context of hybridization and other confounding processes. Various international partners are involved, and I will set up a close collaboration between the University of Antwerp and PUCRS, a Brazilian university with expertise in Neotropical carnivores. Part of the data required for this project is already available at PUCRS, where I contributed to the preliminary results that guide the two objectives of this project: (1) use complete genomes, including from museum specimens, to clarify the evolutionary relationships between the different species of small spotted cats in Latin America, and (2) complement the first objective with novel methods based on alternative genomic markers and partitions to maximize the amount of information that can be gained from these genomic sequences.
Team: Jonas Lescroart, Hannes Svardal
Collaborators: Eduardo Eizirik (Pontifical Catholic University of Rio Grande do Sul)
Funding: FWO PhD Fellowship
Does hybridization facilitate explosive speciation of Lake Baikal amphipods?

The adaptive radiation of amphipods in Lake Baikal has brought forward more than 340 species (20% of the world’s freshwater amphipods), making it one of the largest species flocks after the famous African cichlid radiations, and one of the only large radiations in temperate climates. Despite this iconic status the radiation has not yet been subject to detailed genomic investigation. In this project, we use genomic approaches to test whether ancient hybridization may have facilitated explosive speciation of the group. Hybridization role in emergence of big species flocks is widely discussed: it could be source of additional genetic variation for new adaptations, and also it causes Dobzhansky-Muller incompatibilities which may help to form interspecies barriers. However, it’s kind of chicken and egg problem: there is almost no proof that hybridization is a real cause of adaptive radiation, rather than a consequence of it. Our goal is to construct genome-based alignments of representative species of the group (at the photo), and to use diverse evolutionary genomics techniques to study the history and driving forces of Lake Baikal Amphipoda speciation. Please, feel free to contact us if you are interested in collaboration.