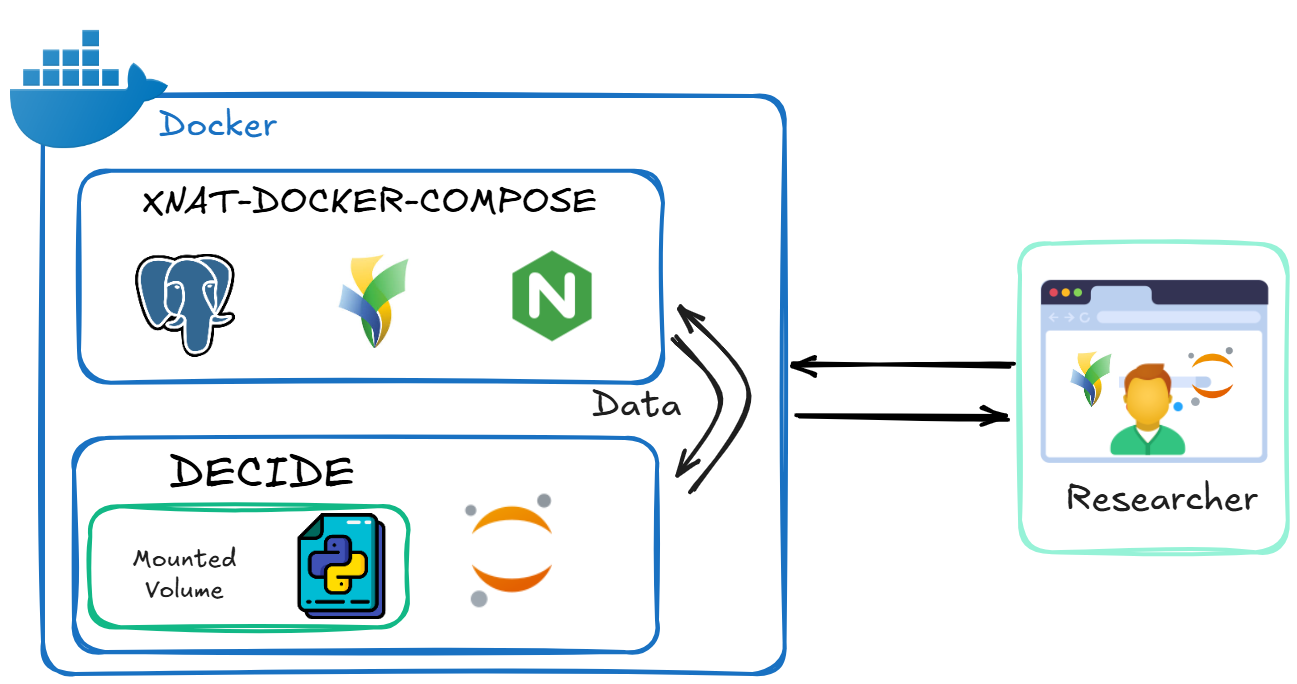
This repository contains scripts and configuration files to build and run a Docker container for image segmentation, post-editing, and (soon) anomaly detection as part of the DECIDE project.
NB: If you are using cloud computing and port forwarding, please refer to:
Cloud Compute Deployment (In ARGOS-1 directory for XNAT) for additional setup instructions.
- Clone the repository to your machine:
git clone https://github.com/NKI-RT/DecideDocker.git
- Navigate into the directory:
cd DecideDocker - Configure environment variables:
- Edit the
workspace/.envfile (make sure hidden files are visible). - Or, rename
default.envto.env.
- Edit the
Start the container using Docker Compose:
sudo docker compose up -d🔄 Note:
The container will perform runtime installations and builds via thestartup.shscript. This may take a few minutes.
- URL:
Openhttp://<your_ip_address>:8888in your browser. - Login Token:
Use the configured token (default:token123).
To stop the container:
sudo docker compose downFor environments using the deprecated standalone docker-compose tool, use:
Start:
sudo docker-compose -f docker-compose-legacy.yml up -dStop:
sudo docker-compose -f docker-compose-legacy.yml downIf you're using cloud computing and your virtual machine's IP address is not public, follow these steps to deploy XNAT and Decide together using a shared proxy network.
-
Stop the existing XNAT container (of ARGOS-1):
# in ARGOS-1 cd xnat-docker-compose
# stop the XNAT sudo docker compose down -
Download this repository into the same directory.:
git clone https://github.com/NKI-RT/DecideDocker.git
-
Copy the cloud compute Docker Compose file:
cp DecideDocker/docker-compose-cloudcompute.yml ./
-
Run the integrated deployment:
sudo docker compose -f docker-compose-cloudcompute.yml up -d
-
To stop the deployment:
sudo docker compose -f docker-compose-cloudcompute.yml down
Congratulations! You can now access your XNAT at:
http://xnat-nginx:80
- This setup runs XNAT, Decide, and NGINX in a shared
proxynetDocker network. - The container will expose:
- JupyterLab (Decide) on port
8888 - XNAT NGINX proxy on port
80
- JupyterLab (Decide) on port
To update your local copy with the latest changes from the original repository and discard any local changes:
-
Add the original repo as a remote (if not already):
git remote add upstream https://github.com/NKI-RT/DecideDocker.git
-
Fetch the latest changes from upstream:
git fetch upstream
-
Reset your local
mainbranch to matchupstream/main:git checkout main git reset --hard upstream/main
-
(Optional) Clean up untracked files and directories:
git clean -fd
⚠️ Warning: This will permanently discard all local changes, including uncommitted edits and untracked files.
We will release more features and scripts/notebooks as the project progresses.
Stay tuned and keep your local copy up to date!
DecideDocker/
├── Dockerfile
├── docker-compose.yml
├── docker-compose-legacy.yml
├── docker-compose-cloudcompute.yml
├── startup.sh
├── workspace/
│ ├── .env (default.env)
│ ├── decide/
│ │ ├── config/
│ │ │ ├── config_total_segmentator.yaml
│ │ │ └── dicomdata_config.yaml
│ │ ├── logs/
│ │ ├── data/
│ │ | └── LUNG1-001/
│ │ | ├── CT
│ │ | ├── RTSTRUCT
│ │ | └── NIfTI
│ │ ├── src/
│ │ │ └── decide/
│ │ └── pyproject.toml
│ ├── notebooks/
│ │ ├── 001_test_essentials.ipynb
│ │ ├── 002_get_nifti.ipynb
│ | └── 002_get_niftiv2.ipynb
│ └── scripts/
│ └── get_nifti.py
└── nnUNet/
The Dockerfile uses a multi-stage build:
- Base:
ubuntu:22.04 - Installs latest JupyterLab and extensions
- Base: TotalSegmentator image
- Adds JupyterLab from Stage 1
- Installs:
- Plastimatch
- PyRadiomics
- nnUNet patch for Plastimatch
- DECIDE tools (including XNAT Python API)
- Resolves dependency issues
- Launches JupyterLab on port
8888
Defines container behavior:
- Uses host IPC (required for nnUNet models)
- Enables GPU support
- Mounts
workspace/directory for live sync between host and container
The startup.sh script is executed when the container starts. It performs the following tasks:
- Builds and installs Plastimatch from source
- Installs required Python packages (e.g., PyRadiomics, Platipy)
- Applies a patch to nnUNet for compatibility
- Installs DECIDE tools if present in
/workspace/decide - Fixes specific package versions (e.g.,
numpy,pydicom) - Registers the Jupyter kernel
- Configures and launches JupyterLab on port
8888
Stores environment variables:
JUPYTER_TOKEN=token123- Mounted inside the container
- Contains all scripts, notebooks, and configurations
src/includes Python packages for segmentation, post-editing, etc.
When installed platipy platipy[nnunet] or platipy[cardiac] and deployed, encountered the error:
UnpicklingError: Weights only load failed...
PyTorch 2.6 changed the default weights_only argument in torch.load to True.
-
Use nnUNet version
1.7.0:git clone https://github.com/MIC-DKFZ/nnUNet.git cd nnUNet git checkout tags/v1.7.0 -b nnunet-1.7.0 -
Patch
model_restore.py:from torch.serialization import safe_globals import numpy with safe_globals([numpy.dtype]): all_params = [torch.load(i, map_location=torch.device('cpu'), weights_only=False) for i in all_best_model_files]
-
Single Docker Compose managing XNAT, JupyterLab, and tools
-
Adding new containers required editing XNAT's Docker Compose
-
Separation of data containers and tool containers
-
No modification needed to data containers when adding/removing tools
This repository includes data derived from the NSCLC-Radiomics dataset available via the Imaging Data Commons (IDC).
Original dataset citation: The Cancer Imaging Archive. https://www.cancerimagingarchive.net/collection/nsclc-radiomics/
License: CC BY-NC 3.0 — Non-commercial use only.