This project aims to identify research articles which are relevant to the Neotoma Paleoecological Database (Neotoma), extract data relevant to Neotoma from the article, and provide a mechanism for the data to be reviewed by Neotoma data stewards then submitted to Neotoma. It is being completed as part of the University of British Columbia (UBC) Masters of Data Science (MDS) program in partnership with the Neotoma Paleoecological Database.
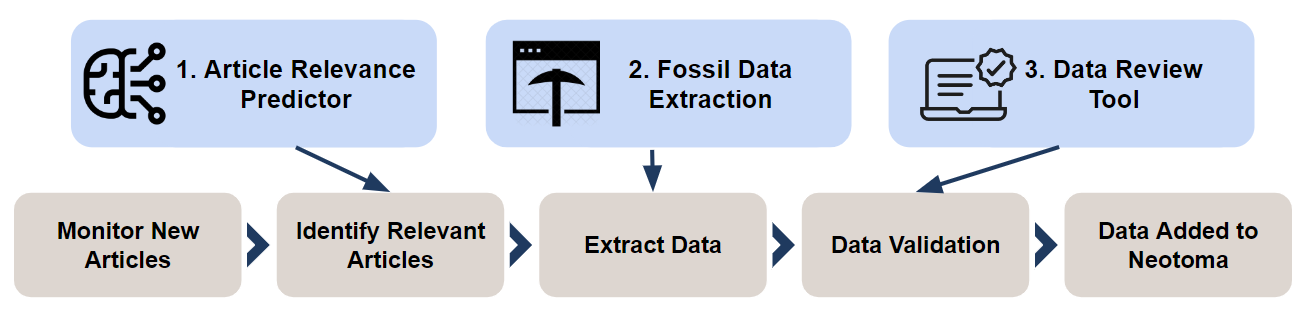
There are 3 primary components to this project:
- Article Relevance Prediction - get the latest articles published, predict which ones are relevant to Neotoma and submit for processing.
- MetaData Extraction Pipeline - extract relevant entities from the article including geographic locations, taxa, etc.
- Data Review Tool - this takes the extracted data and allows the user to review and correct it for submission to Neotoma.
The goal of this component is to monitor and identify new articles that are relevant to Neotoma. This is done by using the public xDD API to regularly get recently published articles. Article metadata is queried from the CrossRef API to obtain data such as journal name, title, abstract and more. The article metadata is then used to predict whether the article is relevant to Neotoma or not. The predicted articles are then submitted to the Data Extraction Pipeline for processing.
The full text is provided by the xDD team for the articles that are deemed to be relevant and a custom trained Named Entity Recognition (NER) model is used to extract entities of interest from the article.
The entities extracted by this model are:
- SITE: name of the excavation site
- REGION: more general regions names to provide context for where sites are located
- TAXA: plant or animal fossil names
- AGE: historical age of the fossils, eg. 1234 AD, 4567 BP
- GEOG: geographic coordinates indicating the location of the site, eg. 12'34"N 34'23"W
- EMAIL: researcher emails referenced in the articles
- ALTI: altitudes of sites, eg. 123 m a.s.l (above sea level)
The model was trained on ~40 existing Paleoecology articles manually annotated by the team consisting of ~60,000 tokens with ~4,500 tagged entities.
The trained model is available for inference and further development on huggingface.co here.

Finally, the extracted data is loaded into the Data Review Tool where members of the Neotoma community can review the data and make any corrections necessary before submitting to Neotoma. The Data Review Tool is a web application built using the Plotly Dash framework. The tool allows users to view the extracted data, make corrections, and submit the data to be entered into Neotoma.
WIP
WIP
WIP
WIP
WIP
├── .github/ <- Directory for GitHub files
│ ├── workflows/ <- Directory for workflows
├── assets/ <- Directory for assets
├── data/ <- Directory for data
│ ├── entity-extraction/ <- Directory for named entity extraction data
│ │ ├── raw/ <- Raw unprocessed data
│ │ ├── processed/ <- Processed data
│ │ └── interim/ <- Temporary data location
│ ├── article-relevance/ <- Directory for data related to article relevance prediction
│ │ ├── raw/ <- Raw unprocessed data
│ │ ├── processed/ <- Processed data
│ │ └── interim/ <- Temporary data location
│ ├── data-review-tool/ <- Directory for data related to data review tool
│ │ ├── raw/ <- Raw unprocessed data
│ │ ├── processed/ <- Processed data
│ │ └── interim/ <- Temporary data location
├── results/ <- Directory for results
│ ├── article-relevance/ <- Directory for results related to article relevance prediction
│ ├── ner/ <- Directory for results related to named entity recognition
│ └── data-review-tool/ <- Directory for results related to data review tool
├── models/ <- Directory for models
│ ├── entity-extraction/ <- Directory for named entity recognition models
│ ├── article-relevance/ <- Directory for article relevance prediction models
├── notebooks/ <- Directory for notebooks
├── src/ <- Directory for source code
│ ├── entity_extraction/ <- Directory for named entity recognition code
│ ├── article_relevance/ <- Directory for article relevance prediction code
│ └── data_review_tool/ <- Directory for data review tool code
├── reports/ <- Directory for reports
├── tests/ <- Directory for tests
├── Makefile <- Makefile with commands to perform analysis
└── README.md <- The top-level README for developers using this project.
This project is an open project, and contributions are welcome from any individual. All contributors to this project are bound by a code of conduct. Please review and follow this code of conduct as part of your contribution.
The UBC MDS project team consists of:
- Ty Andrews
- Kelly Wu
- Jenit Jain
- Shaun Hutchinson
Sponsors from Neotoma supporting the project are:
Issues and bug reports are always welcome. Code clean-up, and feature additions can be done either through pull requests to project forks or project branches.
All products of the Neotoma Paleoecology Database are licensed under an MIT License unless otherwise noted.