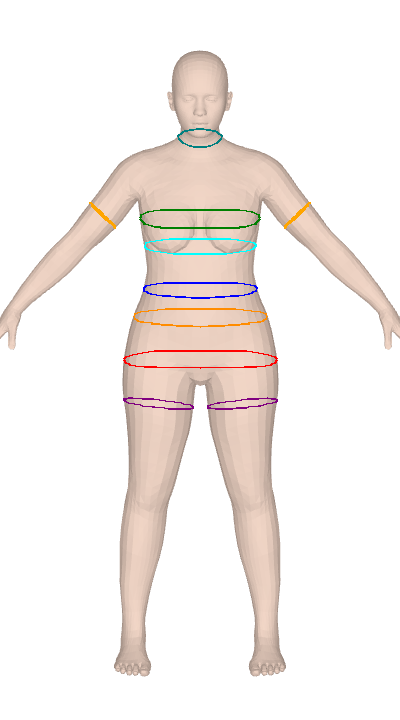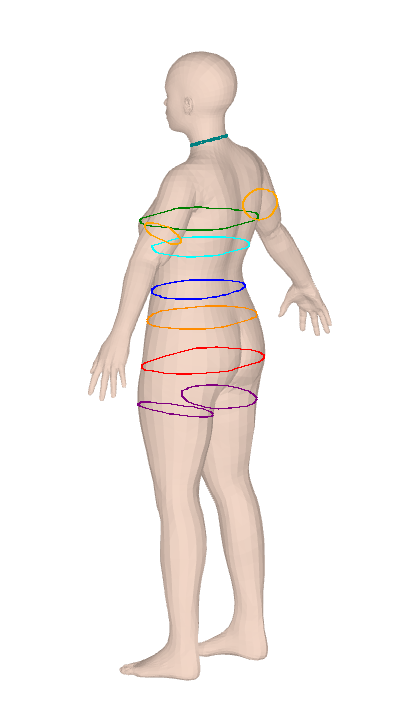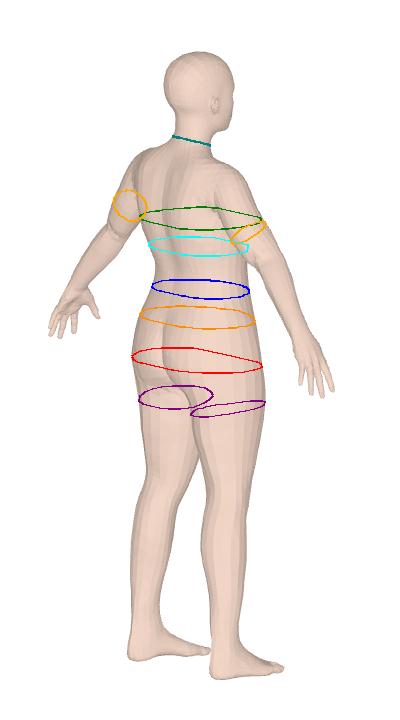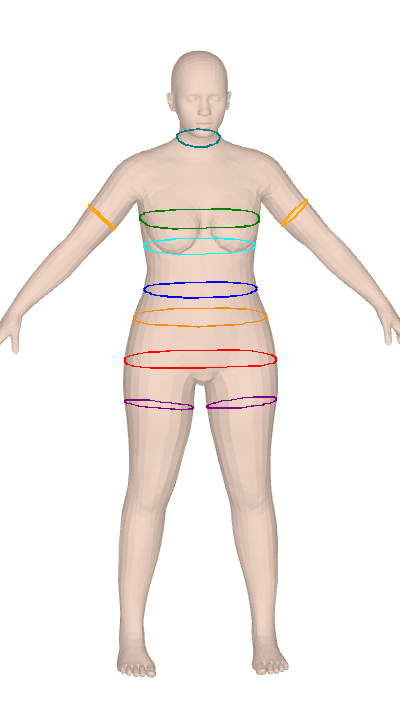
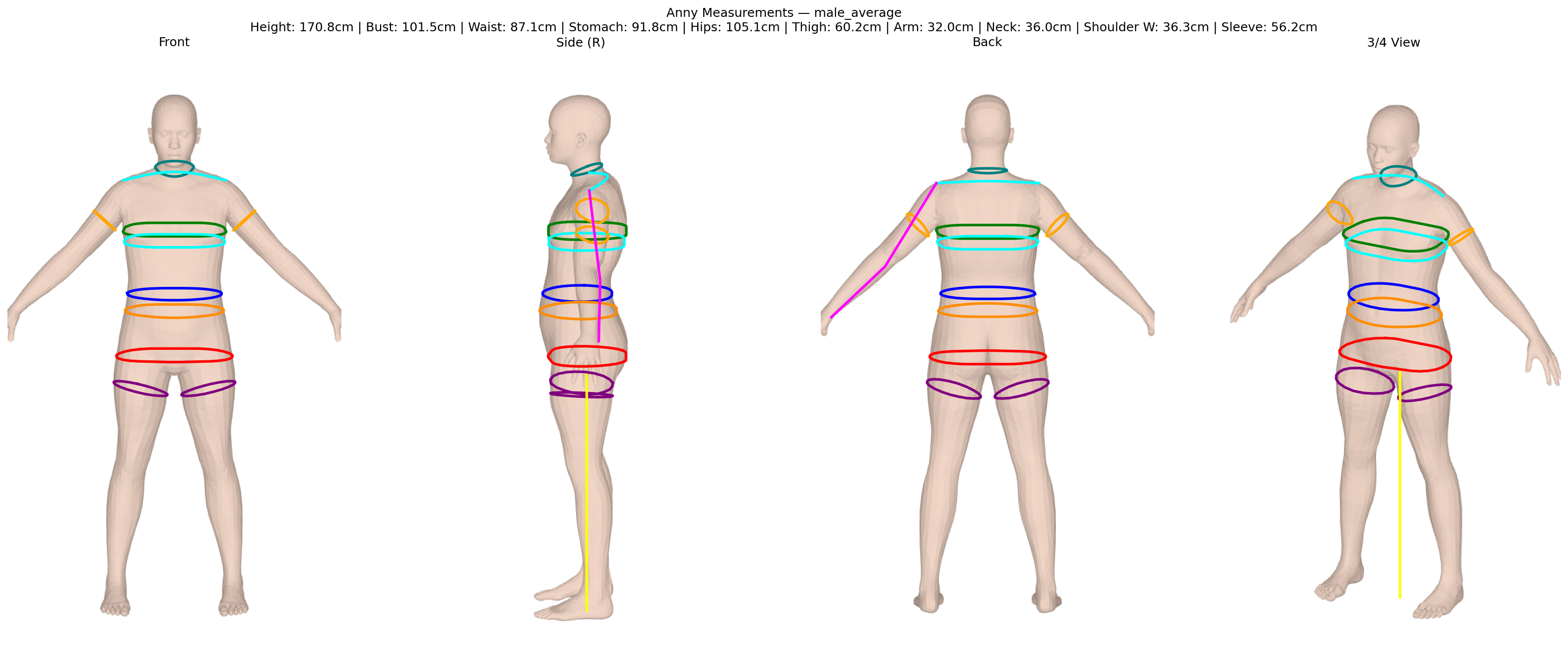
ISO 8559-1 body measurements for Anny and MHR parametric body models. Twelve keys are differentiable through PyTorch autograd for gradient-based body fitting.
Anny and MHR give you a 14–18K vertex mesh and nothing to measure it with. SMPL tooling doesn't port over, and the plane-sweep algorithms look simple until you hit convex-hull tape simulation, contour-fragment merging, and ISO-compliant landmark detection for bust/hip/crotch. clad-body is that work, done once — 25 anthropometric measurements over circumferences, lengths, and body composition (volume, mass, BMI, body fat), calibrated against real scan data. It's used in production at Clad for size-aware virtual try-on.
Prefer REST? Same engine is available as a hosted API at api.clad.you — questionnaire or photo → GLB mesh + the ISO measurements below. Free tier (5 calls/week), grab a key at clad.you/developers. Good fit if you want the body model without the Python install or the Anny model weights.
All bodies are normalised to the same coordinate convention: Z-up, metres, XY-centred, feet at Z=0, +Y=front.
pip install clad-body
# With Anny body loader (requires torch)
pip install 'clad-body[anny]'
# With MHR body loader (requires pymomentum)
pip install 'clad-body[mhr]'
# With 4-view rendering
pip install 'clad-body[render]'from clad_body.load import load_anny_from_params
from clad_body.measure import measure
body = load_anny_from_params(params)
m = measure(body) # all measurements
m = measure(body, preset="core") # 4: height, bust, waist, hip
m = measure(body, preset="standard") # 9: + thigh, upperarm, shoulder, sleeve, inseam
m = measure(body, preset="tops") # garment-relevant subset
m = measure(body, only=["bust_cm", "hip_cm"]) # specific keys
m = measure(body, tags={"type": "circumference", "region": "leg"}) # tag filter
m = measure(body, render_path="body.png") # with 4-view renderMHR works the same way:
from clad_body.load import load_mhr_from_params
body = load_mhr_from_params("path/to/sam3d_params.json")
m = measure(body)Under active development. API surface and supported keys may change between minor versions. Twelve keys are differentiable today; more will follow.
For autograd-based optimization of the body mesh, use measure_grad(body) instead of measure(body). Same input, same key names — but the returned values are PyTorch tensors with autograd history, so you can put them directly into a loss and backprop into the Anny phenotype parameters.
Pass requires_grad=True to load_anny_from_params to create the body with gradient-enabled phenotype tensors (stored on body.phenotype_kwargs):
import torch
from clad_body.load import load_anny_from_params
from clad_body.measure import measure_grad
body = load_anny_from_params(initial_params, requires_grad=True)
optimizer = torch.optim.Adam(list(body.phenotype_kwargs.values()), lr=0.01)
for step in range(500):
optimizer.zero_grad()
m = measure_grad(body, only=["bust_cm", "waist_cm", "inseam_cm"])
loss = (m["bust_cm"] - 92.0) ** 2 + (m["waist_cm"] - 78.0) ** 2 + (m["inseam_cm"] - 82.0) ** 2
loss.backward()
optimizer.step()Each measure_grad(body) call re-runs the forward pass using body.phenotype_kwargs, so after optimizer.step() updates the tensors the next iteration measures the new mesh. If you only want to optimize a subset of parameters, load without requires_grad=True and enable it per-tensor: body.phenotype_kwargs["height"].requires_grad_(True).
Supported keys and their calibration error vs the ISO reference that measure() uses:
| Key | Error vs ISO |
|---|---|
height_cm, waist_cm |
exact (same loop / extent) |
bust_cm |
MAE 0.06 cm, max 0.18 cm |
underbust_cm |
MAE 0.39 cm, max 1.61 cm |
hip_cm |
MAE 0.46 cm, max 1.39 cm |
stomach_cm |
MAE 0.93 cm, P95 2.83 cm, max 3.61 cm (soft-argmin picks a different Z than the reference's 2 mm band scan; residual is inherent Z-choice noise) |
inseam_cm |
RMS 0.06 cm, max 0.10 cm |
sleeve_length_cm |
RMS 0.33 cm, max 0.55 cm |
shoulder_width_cm |
RMS 1.39 cm on 100 random bodies (91 % within ±2 cm). Max 5cm |
upperarm_cm |
≤ 1 cm |
mass_kg |
≤ 3 kg |
thigh_cm |
MAE 0.06 cm, max 0.18 cm (100 random bodies) |
Both measure() and measure_grad report convex hull circumference, not the raw cross-section perimeter. This matches ISO 8559-1: a real measuring tape bridges across concavities (e.g., cleavage between breasts, armpit crease) rather than dipping into them. The convex hull perimeter is always ≤ the raw contour perimeter — the difference is most visible at the bust on larger cup sizes where the cleavage concavity can shorten the measurement by 1-3 cm compared to following the actual surface.
measure_grad builds a 72-point polygon via differentiable soft edge-plane intersection, then takes its scipy.spatial.ConvexHull perimeter. This is used for bust, underbust, hip, and stomach — each at its respective anatomical height. Every measure_grad() call recomputes the hull from scratch (new forward pass → new polygon → new hull). The hull decides which polygon vertices to keep (discrete, like argmax or tensor[mask]) — but the perimeter of those vertices is a plain sum of torch.linalg.norm over their positions, so loss.backward() flows gradients through the kept vertices back to the Anny phenotype params. Dropped vertices (inside the hull, e.g., cleavage or gluteal cleft bins) get zero gradient — correct, since they don't affect the tape measure. The hull indices can change between optimization steps if the body shape changes enough, but the perimeter value is continuous at those transitions so the loss doesn't jump. See clad_body/measure/_soft_circ.py.
For stomach_cm the cutting-plane height is itself differentiable: a soft-argmin over torso vertex Y (most anterior = most negative) in the belly Z-range picks the height of maximum anterior protrusion, and one soft_circumference is taken there. See measure_stomach_soft in the same file. The Z selection can drift by 1–2 cm from the reference's 2 mm band-scan argmax on bodies where multiple vertex clusters have near-identical anterior Y — which translates to ~3 cm of circumference error because the body tapers rapidly in the belly region. This is why stomach_cm has looser tolerance than the other soft-circ keys.
Strictly speaking, stomach_cm is differentiable almost everywhere, with numerically-tiny (~10⁻⁵ cm) step discontinuities where a vertex crosses the hard Z-mask boundary that gates out feet and bust from the belly soft-argmin.
Requesting any other key raises ValueError. There is no silent numpy fallback — it would break gradient flow without warning. For non-differentiable keys use measure().
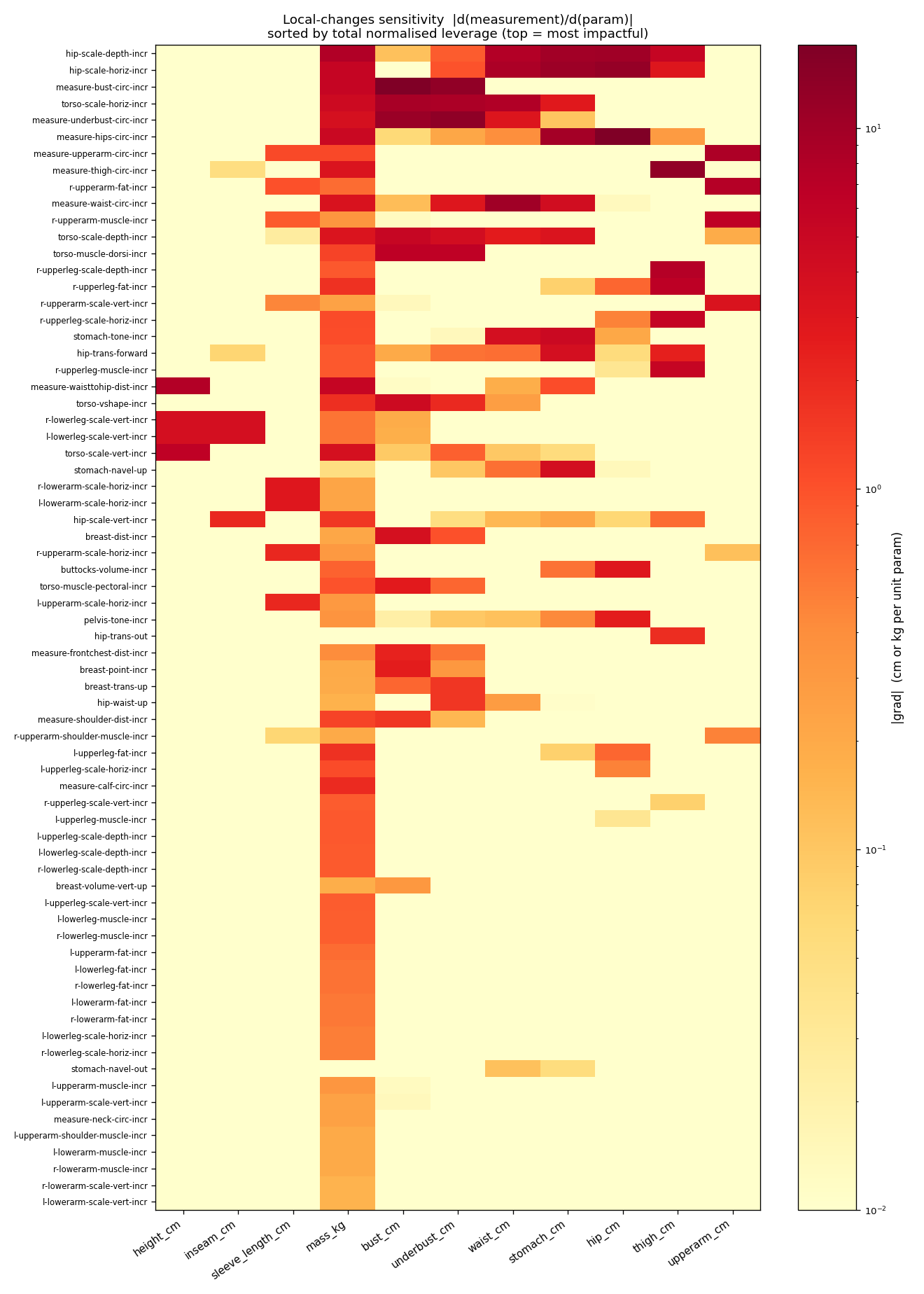
Jacobian heatmap for every Anny local_changes dimension against every supported measurement — |d(measurement)/d(param)| averaged across 6 reference bodies. Darker cells mean the local_change has strong leverage over that measurement; pale cells mean the gradient is near zero and Adam will barely move it. Useful when deciding which params to unfreeze for a given fit target (e.g. optimising bust_cm is pointless without measure-bust-circ-incr or torso-scale-horiz-incr in the active set). Regenerate with python tools/sensitivity_map.py --output <path>.
| Import | What |
|---|---|
clad_body.load.load_anny_from_params |
Load Anny body from phenotype params |
clad_body.load.load_mhr_from_params |
Load MHR body from SAM 3D Body params |
clad_body.load.AnnyBody, MhrBody |
Body dataclasses |
clad_body.measure.measure |
Measure a body (numpy reporting path, ISO 8559-1) |
clad_body.measure.measure_grad |
Differentiable measurements for autograd loops (Anny only) |
clad_body.measure.REGISTRY |
All measurement definitions (dict[str, MeasurementDef]) |
clad_body.measure.list_measurements |
Query measurements by tags |
clad_body.measure.MeasurementDef |
Measurement definition type |
measure() accepts preset, only, tags, exclude. Precedence: only > preset > tags > default ("all"). exclude is applied last. Only runs computation groups needed for the requested keys.
from clad_body.measure import REGISTRY, list_measurements
REGISTRY["bust_cm"].description # self-measurement instructions
REGISTRY["bust_cm"].iso_ref # "5.3.4"
REGISTRY["bust_cm"].type # "circumference"
list_measurements(type="circumference", region="leg") # [thigh, knee, calf]Every measurement is tagged across 5 dimensions. Each carries a human-readable description for self-measurement instructions and i18n key mapping.
| Dimension | Values |
|---|---|
| type | circumference, length, scalar |
| standard | iso (ISO 8559-1), tailor (industry standard), derived (computed) |
| region | neck, torso, abdomen, arm, leg, full_body |
| tier | core > standard > enhanced > fitted (cumulative) |
| garments | tops, bottoms, dresses, outerwear, underwear |
| Preset | Count | Adds |
|---|---|---|
core |
4 | height, bust, waist, hip |
standard |
9 | thigh, upperarm, shoulder_width, sleeve_length, inseam |
enhanced |
18 | neck, underbust, stomach, mass, volume, bmi, body_fat, belly_depth, back_neck_to_waist |
fitted/all |
25 | knee, calf, wrist, crotch_length, front_rise, back_rise, shirt_length |
Garment codes: Tops, Bottoms, Dresses, Outerwear, Underwear.
Tier codes: core, std (standard), enh (enhanced), fit (fitted). Anny-only: underbust, mass, volume, bmi, body_fat, belly_depth.
| Group | Measurements | Cost | Deps |
|---|---|---|---|
| A Core torso | height, bust, waist, hip, stomach, underbust, belly_depth | Cheap | -- |
| B Limb sweeps | thigh, knee, calf, upperarm | Expensive | -- |
| C Joint linear | shoulder_width, sleeve_length (ISO surface walk on re-posed body) | Very expensive | -- |
| D Perpendicular | neck, wrist | Medium | -- |
| E Mesh geometry | inseam (mesh sweep), crotch_length, front_rise, back_rise | Medium | -- |
| F Surface trace | shirt_length | Medium | E |
| G Body composition | volume, mass, bmi, body_fat | Cheap | D |
| H Back length | back_neck_to_waist | Cheap | A |
Group C and E have differentiable alternatives in measure_grad — use it for hot-loop optimization instead of calling measure() repeatedly.
measure() only runs the computation groups needed for the requested keys — use only= or preset= to skip expensive groups:
measure(body) # all groups — ~800 ms
measure(body, preset="core") # group A only — ~100 ms
measure(body, only=["bust_cm"]) # group A only — ~100 ms
measure(body, only=["shoulder_width_cm"]) # groups A + C — ~200 msmeasure() accepts a device parameter (None = auto-detect CUDA):
measure(body, only=["bust_cm"], device="cuda") # GPU forward pass
measure(body, device=None) # auto: CUDA if available| Extra | What it enables |
|---|---|
[anny] |
Anny body loader (requires torch) |
[mhr] |
MHR body loader (requires pymomentum) |
[render] |
4-view body renders (requires matplotlib, pyrender) |
Without extras, only numpy, scipy, and trimesh are required.
- Consumer demo — full pipeline (questionnaire → body → virtual try-on) at clad.you/size-aware/size-me.
- REST API — same body model + these measurements behind a bearer-token API at api.clad.you. Swagger UI on the root; 5 calls/week free tier. Key management at clad.you/developers.
This library was built for Clad's size-aware virtual try-on pipeline. Read the full story: A 3D Body Scan for Nine Cents — Without SMPL.
Apache 2.0 — see LICENSE.